Advances in optical microscopy and molecule-specific fluorescence labelling strategies have enabled molecular insights into the spatiotemporal dynamics of biological processes that were previously inaccessible. In the Laboratory for Cellular Biophysics, we specialise in parallel measurement of multiple fluorescence observables – intensity, anisotropy, and lifetime – to quantify molecular behaviour in the native environment of the living nucleus. By coupling these observables with FRET and fluorescence correlation spectroscopy (FCS), we can resolve: (1) structure at the nanometre scale, using FRET to detect protein–protein and protein–ligand interactions, and (2) dynamics across nanoseconds to milliseconds, using FCS and related approaches to quantify diffusion, transport, and interaction kinetics. This methodological framework enables us to measure key molecular parameters, including protein–ligand binding, oligomerisation, and intracellular trafficking.
Core methods:
FLIM Fluorescence lifetime imaging microscopy: used for hetero-FRET detection and mapping of protein interactions, as well as biosensing of intracellular pH.
FAIM Fluorescence anisotropy imaging microscopy: used for homo-FRET detection and mapping of protein self-association, as well as biosensing of intracellular viscosity.
FCS Fluorescence correlation spectroscopy: used to quantify local protein dynamics in a subcellular location (e.g. diffusion coefficient and concentration).
FFS Fluorescence fluctuation spectroscopy and brightness analysis: used to quantify and spatially map protein oligomer formation and stoichiometry.
sFCS, pCF Scanning FCS and pair correlation function analysis: used to quantify protein transport with respect to subcellular architecture (e.g. arrival time, directionality, diffusion law).
STICS, RICS Spatiotemporal image correlation spectroscopy: used to map protein mobility and binding dynamics across a subcellular location.
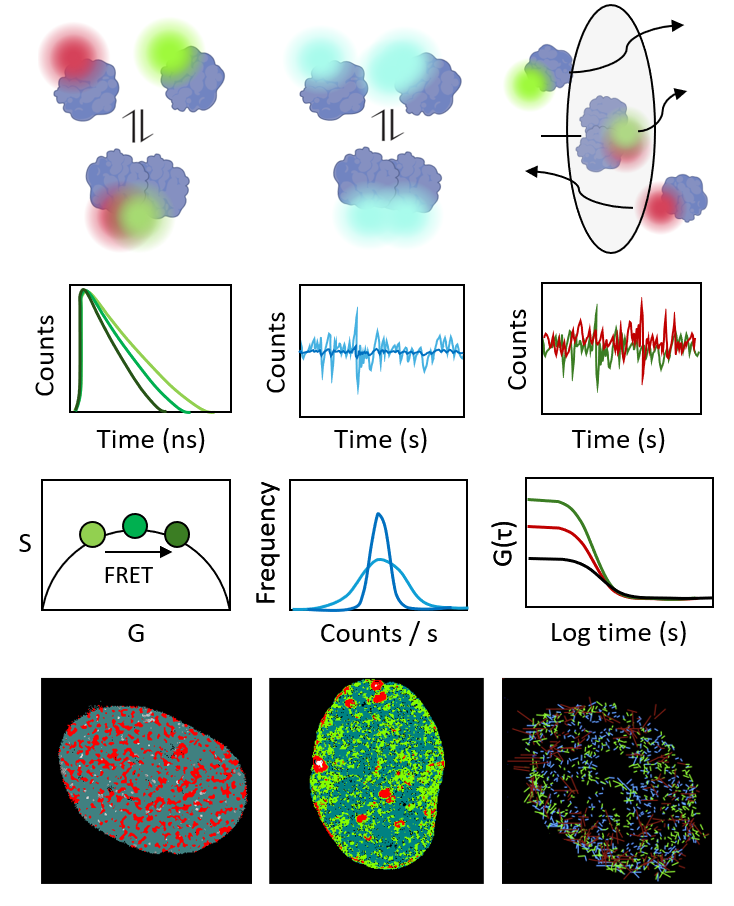
Together, these methods provide a powerful framework to reveal how molecular interactions and dynamics underpin genome function within the context of live-cell nuclear architecture.
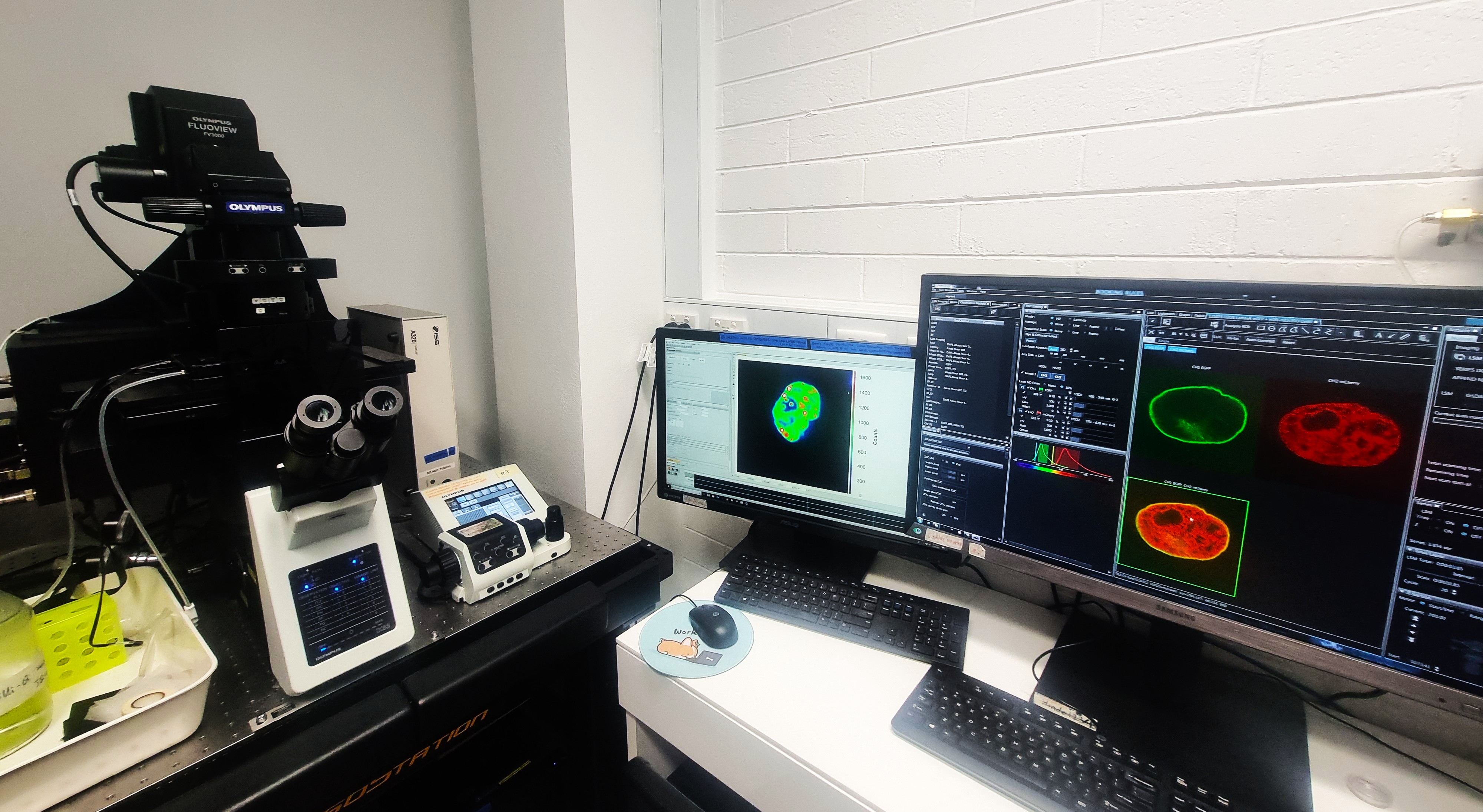
Olympus FV3000
High-performance confocal laser scanning microscope coupled to an ISS FLIMbox, enabling time-resolved fluorescence lifetime detection alongside fluorescence fluctuation spectroscopy and orbital tracking for quantitative live-cell imaging.
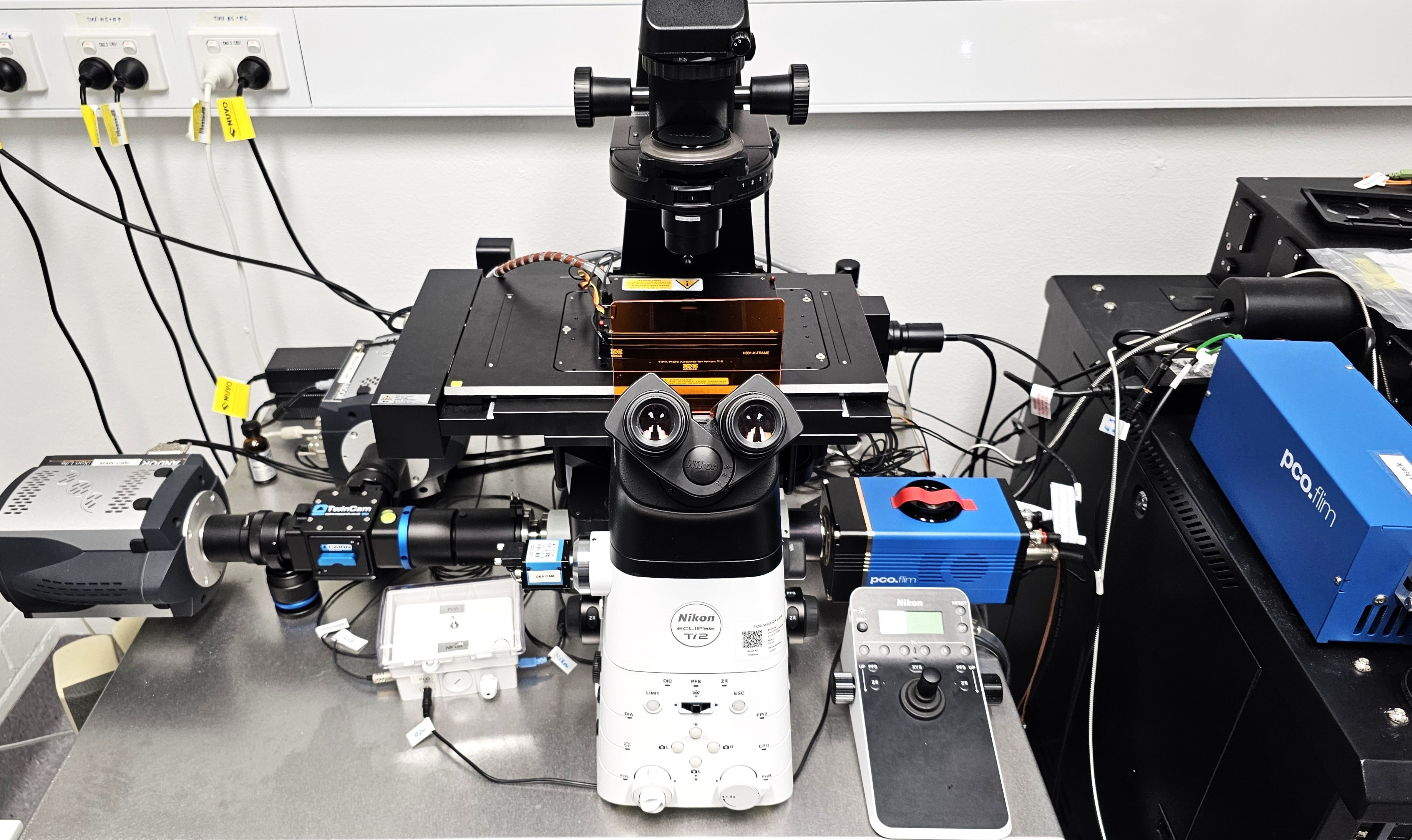

Nikon Ti2 Eclipse
High-performance TIRF and widefield fluorescence microscope coupled to a time-resolved PCO camera and pulsed laser excitation, enabling lifetime-based FRET detection and single-molecule tracking for quantitative live-cell imaging.
For more information visit: microscopy.unimelb.edu.au/om/capabilities/instruments

Parkville VIC 3010
University of Melbourne
Australia
