Architectural organisation of the cell nucleus
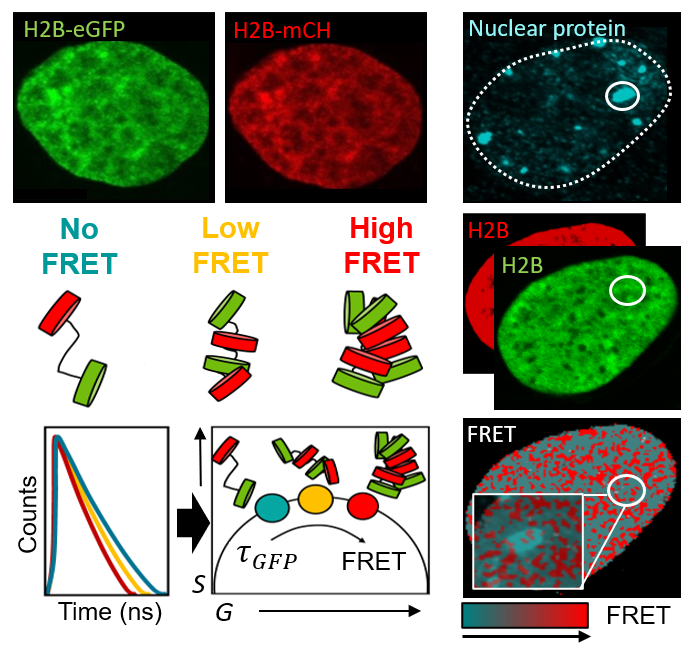
We aim to understand how the two metres of DNA that underpin the human genome are packaged into a 3D chromatin network that fits within the tiny volume of a cell nucleus, and how sub-micron to nanoscale rearrangements in this framework regulate access to the DNA template. To investigate this, we: (1) spatiotemporally map chromatin accessibility via fluorescence fluctuation spectroscopy (FFS) of inert fluorescent tracers of different sizes, and (2) quantify FRET interactions between fluorescently labelled histones (H2B) and architectural proteins (HP1) using FLIM and FAIM. This allows us to study the role of chromatin structure and dynamics in live-cell genome function down to the level of nucleosome proximity.
Relevant Publications
- Lou J., Deng Q., Zhang X., et al. Heterochromatin protein 1 alpha (HP1α) undergoes a monomer to dimer transition that opens and compacts live cell genome architecture. Nucleic Acids Res. (2024)
- Solano A., Lou J., Scipioni L., et al. Radial pair correlation of molecular brightness fluctuations maps protein diffusion as a function of oligomeric state within live-cell nuclear architecture. Biophysical Journal (2022)
- Lou J., Scipioni L., Wright B.K. & Hinde E. Phasor histone FLIM-FRET microscopy quantifies spatiotemporal rearrangement of chromatin architecture during the DNA damage response. PNAS USA (2019)
Protein Navigation of the Nuclear Landscape
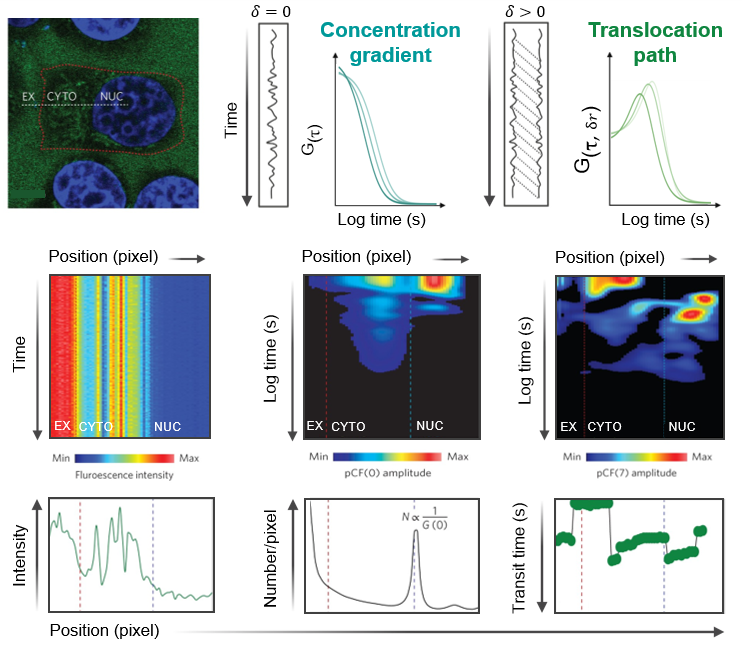
We investigate how DNA-binding proteins navigate the nucleus and how protein–protein interactions regulate DNA target search. To do so, we are developing fluorescence fluctuation-based analyses to extract nuclear trafficking events directly from live-cell microscopy data. Using spatiotemporal correlation functions, we map protein local diffusion to long-range nuclear transport, while resolving how these dynamics shift with oligomeric state or complex formation via use of spectrally distinct fluorescent ligands. These measurements are compatible with single-, dual-, or triple-channel CLSM or TIRF microscopy time series and allow us to determine how DNA repair factors navigate nuclear architecture to coordinate double-strand break resolution, and how transcription factors use protein stoichiometry to regulate DNA target search and gene expression.
Relevant Publications
- Hinde E., Thammasiraphop K., Duong H. et al. Pair correlation microscopy reveals the role of nanoparticle shape in intracellular transport and site of drug release. Nature Nanotech. (2017)
- Lou J., Priest D.G., Solano A., et al. Spatiotemporal dynamics of 53BP1 dimer recruitment to a DNA double strand break. Nat Commun. (2020)
- Sanchez-Velasquez J., Solano A., Digman M.A., et al Pair correlation microscopy of intracellular molecular transport. Nat Protoc. (2025)

Hinde Lab
Laboratory for Cellular Biophysics
David Caro Building
Parkville VIC 3010
University of Melbourne
Australia